The soil bacterium Pseudomonas putida as a non-pathogenic representative of the robust and versatile bacterial family of Pseudomonads is known to be an excellent starting point for the construction of recombinant production strains for rhamnolipid biosurfactants. On the other hand, this bacterium naturally lacks properties to effectively use many biorefinery products for growth and production. So far, plasmids with strong synthetic promoters were overwhelmingly applied for the expression of the rhl operon encoding the production of rhamnolipids; in contrast, genome-integrated expression cassettes are generally considered suitable for the construction of stable strains. Within Bio², the native metabolism of P. putida KT2440 (Chassis - Fig. 1) and the corresponding rhl expression cassette (Cargo - Fig. 2) were addressed in complementary work packages to yield an optimized P. putida cell factory for the production of rhamnolipids.
One aim was the implementation of xylose and ethanol metabolization (Fig. 1 A). The utilization of xylose would enable the use of inexpensive lignocellulosic material and thus increase the added value of a P. putida-based process. To this end, plasmid-based expression of three different bacterial xylose pathways - isomerase, Weimberg and Dahms pathway - was shown to enable xylose-based growth. Expressions of the oxidative Weimberg and Dahms pathways lead to high biomass production. Besides, the initial lag phase could be substantially shortened with the help of adaptive laboratory evolution. Ethanol produced from biomass is another interesting substrate for a biotechnological process due to its direct introduction into the central carbon metabolism as acetyl-CoA without loss of carbon. The natural ability of P. putida to metabolize ethanol has been significantly improved by adaptive laboratory evolution. Both chassis showed was comparable or even better for rhamnolipid production based on the alternative substrates than with the conventional substrate glucose[1],[2]. In addition, the monorhamnolipid titer produced was increased by further genetic optimization of P. putida, e.g. the deletion of energy-intensive processes such as flagellar biosynthesis and activity or competing synthesis pathways such as polyhydroxyalkanoate synthesis[3] (Fig. 1 B). The production was increased even further by an increased supply of rhamnose and fatty acid in activated form as starting materials for rhamnolipid biosynthesis after metabolic engineering.
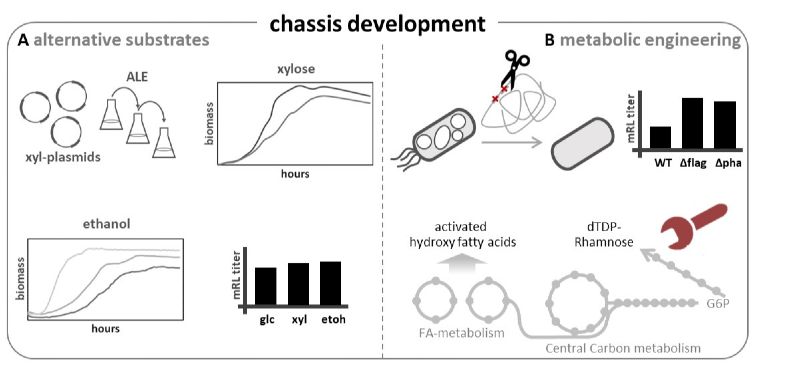
Fig. 1: Chassis development. A – Efficient utilization of the alternative substrates xylose was achieved by using vectors encoding respective catabolic genes in combination with adaptive laboratory evolution (ALE). This resulted in different strains with different growth rates and monorhamnolipid titers on alternative substrates1,2. B – In the course of metabolic engineering, genes for flagella and PHA biosynthesis were deleted, which increased the monorhamnolipid titer3. In addition, targets in the central carbon metabolism were addressed, resulting in an increased supply of activated rhamnose and fatty acid.
P. putida KT2440 does not possess these genes by nature, which makes it possible to regulate their expression as required without interfering with naturally evolved control processes.
As a first step, a new Tn7-transposon was constructed. It combines the stable integration of the biosynthetic gene at a unique site in the genome with the adaptivity of the yTREX toolbox developed in collaboration with the BioSC-FL CombiCom (Fig. 2 A). This new tool enabled the effective construction of a set of strains with new expression cassettes and the subsequent comparative expression studies with different promoters to investigate their influence on gene expression, rhamnolipid titer and strain stability3 (Fig. 2 B). Remarkably, the use of strong, constitutive promoters resulted in the loss of rhamnolipid formation capacity, indicating a negative selection pressure due to rhamnolipid production. In contrast, the inductor-regulated expression systems, here applied for the first time in the context of rhamnolipid biosynthesis, showed efficient target gene expression combined with an outstanding strain stability. Overall, the salicylate-inducible rhlAB expression system led to the best production characteristics. The optimization of the rhamnolipid titer and the complete conversion of the precursor HAA was achieved by its implementation into the genome-reduced strain background and by additional co-integration of relevant educt synthases in the expression cassette (Fig. 2 C). Finally, successful production of different rhamnolipid variants illustrated the broader applicability of this expression system (Fig. 2 D).

Fig. 2: Cargo design. In order to optimize rhamnolipid production by improved expression of the rhamnolipid biosynthesis genes in P. putida, a new integration tool (A) to construct and assess rhamnolipid expression cassettes with different promoters3 (B). The most promising cassette was supplemented with accessory genes to strengthen the metabolic flow towards the product (C). Also, different variants of rhamnolipids could be produced this way (D).
A prerequisite for the parallel consideration of many genetically slightly different expression strains for the optimization of rhamnolipid production was the availability of simple methods that allow a fast and reliable detection, quantification and analysis of the biosurfactants (Fig. 3). Within Bio2, different assays for high-throughput screening have been developed or established for rhamnolipids (Fig. 3 A) including the atomized-oil and the VPBO assay[1] . While such assays can be used to get fast impression of the obtained titers, the suitable analytic platform provided information about the exact amount and composition of the rhamnolipids (Fig. 3 B).
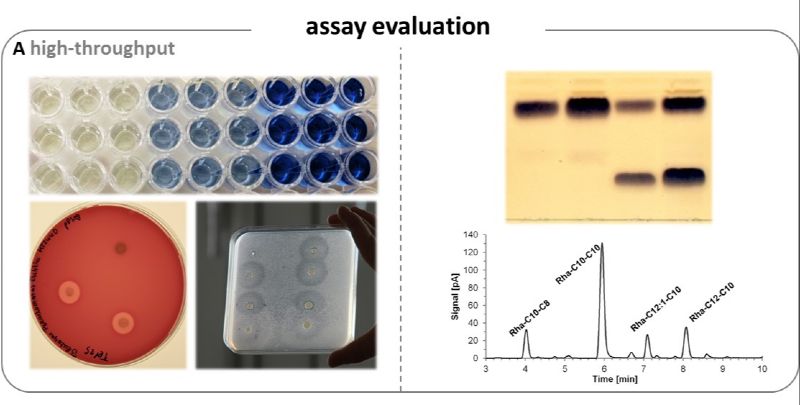
Fig. 3: Detection of rhamnolipids. A - High-throughput screening methods such as VPBO assay[4], blood agar plates and atomized-oil assay facilitate the detection and analysis of suitable producers. B - Analytical detection methods (thin-layer chromatography and high-performance liquid chromatography with Charged Aerosol Detection [HPLC-CAD]) allow the identification and quantification of different rhamnolipid congeners.
[1] https://doi.org/10.3389/fbioe.2019.00480 [2] https://doi.org/10.3389/fbioe.2020.00899
[3] https://doi.org/10.3389/fbioe.2020.00976 [4] https://doi.org/10.3389/fbioe.2020.00958
Prof. Lars Blank
Dr. Till Tiso
Isabel Bator
Institute of Applied Microbiology
RWTH Aachen University
Prof. Karl-Erich Jaeger
Dr. Stephan Thies
Sonja Kubicki
Institute of Molecular Enzyme Technology (IMET)
Heinrich-Heine University Düsseldorf