Pyrrole alkaloids include the family of prodiginines that occur in different bacteria and share common tripyrrole scaffolds with additional variable features, created by a modular biosynthetic pathway. The natural structural differences of prodiginines are known to result in a broad spectrum of different bioactivities like anticancer and antibiotic activity. Therefore, the natural compound family provided an intriguing starting point for the production and investigation of natural and nature-inspired compounds.
We have thus installed prodiginine biosynthesis in the amenable bacterium Pseudomonas putida, which exhibits remarkable tolerance toward the compounds that are toxic to other species. To this end, molecular genetic tools facilitating the transfer of biosynthetic gene clusters and combinatorial gene expression were refined. Here, the ribosomal RNA encoding genes were identified as specifically suitable integration sites for expression of the respective biosynthetic gene cluster, and > 0.2 g/L prodigiosin production was achieved. This could be followed in real-time via newly established online-monitoring of product formation and optimised regarding, among others, the cultivation temperature.
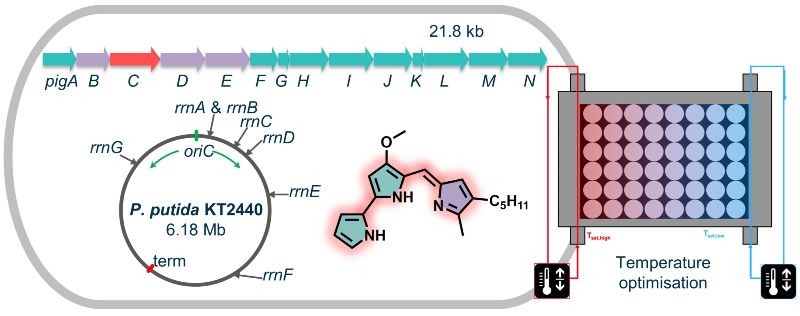
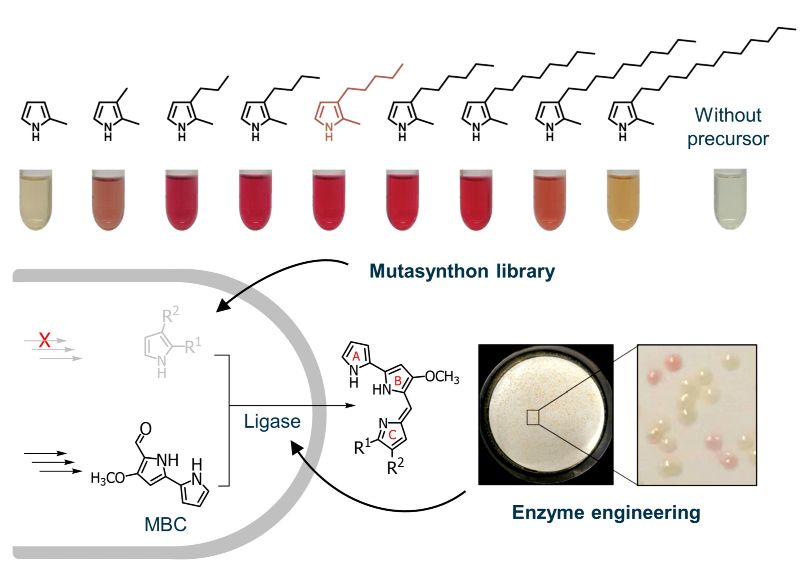
Based on that, we further produced multiple new prodiginines by exploiting the modular prodiginine biosynthesis pathway: Two precursor modules provide a monopyrrole and a bipyrrole precursor, which are fused in the condensation module. This produces the tripyrrolic scaffold conferring the typical red colour to the prodiginines, which can be optionally further processed in a decoration module. This enabled implementation of effective mutasynthesis and semisynthesis concepts for incorporation of externally added precursor analogues, which were obtained by chemical synthesis, into new tripyrroles and product derivatisation. We succeeded to widen the spectrum of prodiginine products by engineering the key ligase enzyme, altogether finally resulting in more than 40 differently substituted prodiginines.
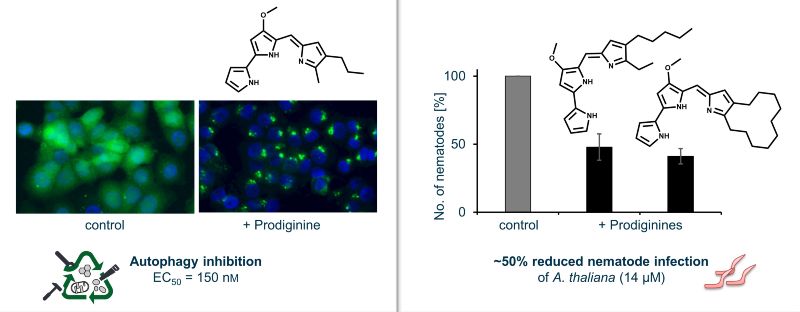
Bioactivity analysis elucidated a range of effects. Some prodiginines showed antibacterial effects and combinatorial activity with surfactants. One of the derivatives showed antiproliferative activity, modulating autophagy and inducing apoptosis in breast cancer cells. Moreover, several prodiginines were active against plant pathogens. Specifically, they inhibited the plant pathogenic fungi Phoma lingam and Sclerotinia sclerotiorum, the causative agents of the blackleg disease of Brassica crops and white mold of diverse host plants including vegetables, rapeseed and ornamentals, respectively. Several prodiginines significantly reduced plant infestation with Heterodera schachtii, a parasitic nematode, which causes severe problems in crops like rapeseed and sugar beet. Some prodiginines even stimulated plant growth. Our interdisciplinary studies demonstrate how the structural modification of a bioactive lead can guide compound activity toward desired properties and hereby introduce new prodiginine compounds that have the potential to be developed into novel plant protection agents.
Our joint efforts thus directly addressed some key challenges of the field. These are the identification of a suitable microbial chassis enabling the efficient production of a compound (family), as well as the effective derivatisation of chemical lead scaffolds. To this end, the necessary synthetic biology tools for biomodule adaptation and engineering were established and implemented. Importantly, the effective combination of synthetic biology and synthetic chemistry was identified as the key synergistic approach to further develop the field.
Prof. Karl-Erich-Jaeger
Dr. Anita Loeschcke
Dr. Thomas Drepper
Robin Weihmann
Fabienne Hilgers
Institute of Molecular Enzyme Technology
HHU Düsseldorf
Prof. Jörg Pietruszka
Dr. Thomas Classen
Jan Gebauer
Hannah Brass
Bioorganic Chemistry
HHU Düsseldorf
Prof. Ulrich Schwaneberg
Dr. Anna Joelle Ruff
Stefanie Brands
Dr. Johannes Schiffels
ABBt – Biotechnology
RWTH University
Prof. Jochen Büchs
Dr. Nina Ihling
Carl Brehl
Biochemical Engineering (AVT.BioVT)
RWTH University
Prof. Florian Grundler
Dr. Sylvia Schleker
Dr. Samer Habash
INRES – Molecular Phytomedicine
University of Bonn