The technology platform BIOExpresSPro provides tools for the identification and production of industry-relevant enzymes that can degrade plant biomass effectively, for example. Using innovative protein-based fluorescence biosensors, secretion processes can now be monitored directly. An automated mini-pilot plant micro bioreactor system with higher process throughput was validated and is now available for strain screening.
An essential part of the bioeconomy is exploiting and utilizing biological resources to produce sustainable and bio-based products. Plants have a key position here since they produce biomass using water, carbon dioxide, and sunlight. They thus represent an almost inexhaustible source of raw material for bioeconomic utilization and industrial value creation. However, to use these plant raw materials sustainably, they must first be made accessible. One strategy to gain this access under environmentally friendly conditions is the application of enzymes.
Hidden champions
Enzymes are biocatalysts that accelerate chemical reactions. In doing so, enzymes are very efficient. An enzyme catalyzes a chemical or biochemical reaction by converting a specific starting molecule, the substrate, into a product. The enzyme itself emerges unaltered from this reaction. Enzymes are thus “hidden champions”, the “workhorses” of industrial biotechnology. Among other things, they are used as helpers in the food and feed industry but also in the cosmetics, textile, paper and pulp industries, the chemical industry, fuel production, the agricultural industry, and the medical and pharmaceutical industries.
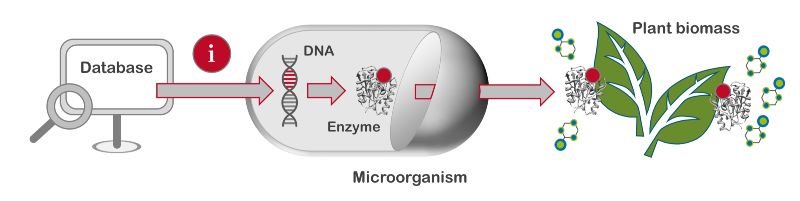
The objective of the BioSC Boost Fund project BIOExpresSPro was to identify in a first step naturally occurring enzymes that degrade plant biomass. Such enzymes can occur in fungi and bacteria, for example, which in nature ensure the decomposition of everything from leaves to entire tree trunks. However, these microorganisms occurring in nature are often not suitable for use in the laboratory. Their demands on the environment, such as nutrient supply and temperature, are usually too complex. Thus, the second objective was to transfer the genes encoding these specialized enzymes into easy-to-handle laboratory organisms, producing and secreting the desired enzymes. In a further step, the enzymes were to be optimized. To achieve these goals, broad expertise from biology, biochemistry, biotechnology, bioprocess engineering, bioanalytics, and bioinformatics was required. Furthermore, BIOExpresSPro has been advised by two associated industry partners throughout the project. These partners were AB Enzymes GmbH and evocatal GmbH, which Advanced Enzyme Technologies Ltd. has since acquired as evoxx technologies GmbH.
Pooling of expertise
As a technology platform, BIOExpresSPro channels this broadly based expertise and industry knowledge into so-called pipelines. Thus, a prediction pipeline was initially established to identify new biomass-degrading enzymes. Using sophisticated algorithms, publicly accessible databases such as BRENDA and UniProt were bioinformatically searched. By doing so, unknown enzymes were found whose features are in great demand also by industry partners. Such enzymes were found in the bacterial genera Arthrobacter and Pseudomonas, for example. To use such enzymes in bioeconomic processes, they must be produced in sufficient quantities. Hence, in another pipeline, the genes of the bioinformatically identified project-relevant enzymes were introduced into easy-to-handle microbial hosts using state-of-the-art cloning methods. These hosts then synthesized the sought-after enzymes in large quantities. By directed evolution of the host cells, the scientists also ensured that the desired enzymes were released in larger quantities from the host cells and did not remain in the cell interior. This so-called secretion facilitates the subsequent harvest of the enzymes immensely.
An extensive genetic library of the enzyme cutinase from the fungus Fusarium solani pisi was generated and used in different host organisms to demonstrate the method’s feasibility. Cutinases can crack the waxy protective armor of plants. They open the gates into the plant and make them accessible for further degradation processes. These enzymes thus play an essential role in the industrial utilization of plant biomass. In a subsequent step, it was possible to optimize cutinase production in Escherichia coli and Corynebacterium glutamicum, which are widely used bacteria in biotechnology.
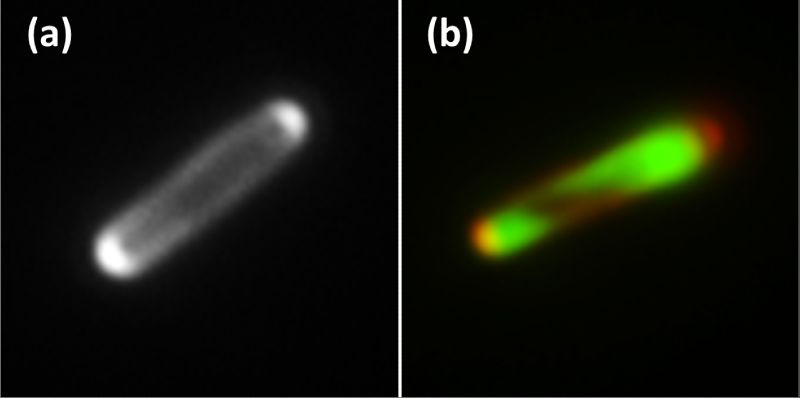
Successful production of enzymes
Scientists of the technology platform BIOExpressSPro also developed novel biosensors to monitor the secretion of the synthesized desired enzyme from the cell in a concentration- and location-dependent manner. The process is based on a fluorescent protein that lights up cells that secrete particularly heavily. Using this technique, it was possible to isolate a new hypersecretion mutant of C. glutamicum that secreted significantly higher quantities of the desired enzyme into the medium than the wild-type strain. In parallel, another novel fluorescent sensor was established, which is used to analyze secretion and protein localization in E. coli. This biosensor allows localizing the selected target protein, for example the desired enzyme, in the cell due to its fluorescence emission. Since this biosensor has a different fluorescence inside the cells than outside, it can also determine the secretion rate of the desired enzyme across the inner cell membrane. In addition, an automated mini-pilot plant micro bioreactor system was developed and validated at the level of process development. This bioreactor system helps to find, isolate and cultivate bacterial strains with the desired features in a short time. The results obtained and the methods newly developed in the framework of BIOExpresSPro were so promising that they had been incorporated into further projects that have been and are being funded by the Federal Government, the state of NRW, and the EU.
Project Coordinator
Dr. Achim Heck
Institute of Molecular Enzyme Technology, HHU Düsseldorf
Email
Participating Core Groups
Prof. Dr. Karl-Erich Jaeger, Dr. Achim Heck, Dr. Thomas Drepper, Dr. Katrin Troost, Andreas Woop
Institute of Molecular Enzyme Technology, HHU Düsseldorf
Prof. Dr. Michael Bott, Prof. Dr. Roland Freudl, Sarah-Kristin Jurischka
IBG-1 Biotechnology, Forschungszentrum Jülich
Prof. Dr. Wolfgang Wiechert, Prof. Dr. Marco Oldiges, Dr. Jannick Kappelmann, Dr. Johannes Hemmerich
IBG-1 Biotechnology, Forschungszentrum Jülich
Prof. Dr. Björn Usadel, Dr. Alexander Vogel
IBG-4 Bioinformatics, Forschungszentrum Jülich
Prof. Dr. Ulrich Schwaneberg, Dr. Anna Joëlle Ruff, Dr. Wei Long
ABBt – Biotechnology, RWTH Aachen
Industry partners (associated partners)
Dr. Christian Degering, Dr. Michael Puls, Dr. Christian Leggewie
evocatal GmbH, Düsseldorf (currently: evoxx technologies)
Dr. Jari Vehmaanperä, Prof. Dr. Karl-Heinz Maurer
AB Enzymes GmbH, Darmstadt
Funding Period
1.1.2014 – 31.12.2016
Funding
BioExpresSPro is part of the NRW-Strategieprojekt BioSC and thus funded by the Ministry of Culture and Science of the German State of North Rhine-Westphalia.
Publications
Freier, L, Hemmerich, J, Schöler, K, Wiechert, W, Oldiges, M and von Lieres, E (2016). Framework for kriging-based iterative experimental analysis and design: Optimization of secretory protein production in corynebacterium glutamicum. Engineering in Life Sciences 16(6): 538-549.
Heck, A and Drepper, T (2017). Engineering photosynthetic α-proteobacteria for the production of recombinant proteins and terpenoids. Modern topics in the phototrophic prokaryotes: Environmental and applied aspects. Hallenbeck, P.C. Cham, Springer International Publishing: 395-425.
Hemmerich, J, Rohe, P, Kleine, B, Jurischka, S, Wiechert, W, Freudl, R and Oldiges, M (2016). Use of a sec signal peptide library from bacillus subtilis for the optimization of cutinase secretion in corynebacterium glutamicum. Microbial Cell Factories 15(1): 208.
Loehrer, M, Vogel, A, Huettel, B, Reinhardt, R, Benes, V, Duplessis, S, Usadel, B and Schaffrath, U (2014). On the current status of phakopsora pachyrhizi genome sequencing. Front Plant Sci 5: 377.
MacKellar, D, Lieber, L, Norman, JS, Bolger, A, Tobin, C, Murray, JW, Oksaksin, M, Chang, RL, Ford, TJ, Nguyen, PQ, Woodward, J, Permingeat, HR, Joshi, NS, Silver, PA, Usadel, B, Rutherford, AW, Friesen, ML and Prell, J (2016). Streptomyces thermoautotrophicus does not fix nitrogen. Scientific Reports 6: 20086.
Torra, J, Burgos-Caminal, A, Endres, S, Wingen, M, Drepper, T, Gensch, T, Ruiz-Gonzalez, R and Nonell, S (2015). Singlet oxygen photosensitisation by the fluorescent protein pp2fbfp l30m, a novel derivative of pseudomonas putida flavin-binding pp2fbfp. Photochem Photobiol Sci 14(2): 280-287.