The BioSC Boost Fund project PectiLyse had the objective to use the fungal model organism Ustilago maydis for the degradation of pectin-rich biomass. Large quantities of organic side streams arise during sugar production from sugar beet. These largely consist of the complex polysaccharide pectin. Currently, the pectin-rich side streams are sold as feed with low added value. The PectiLyse project should establish basic methods to convert these residues into valuable products in an integrated bioprocess in the future.
A consortium of researchers from different disciplines fostered an interdisciplinary approach to the project: microbiologists with expertise in genetic manipulation of the fungus as well as the production of organic acids teamed up with leading bioprocess engineers. This supported a holistic approach in which optimized strains could be efficiently characterized using online analytics.
The fungus Ustilago maydis has some attributes that especially qualifies it for the development of an integrated bioprocess. In contrast to other known pectin-degrading fungi, Ustilago maydis is able to grow filamentously as well as yeast-like. This offers decisive advantages in handling and especially in cultivation. Ustilago maydis infects corn plants and causes the so-called corn smut. However, it is not dangerous for humans. Infected corn cobs are even suitable for human consumption and sold as a delicacy in Mexico. Yeast-like growing laboratory strains are not infectious for plants and are ideal for biotechnology.
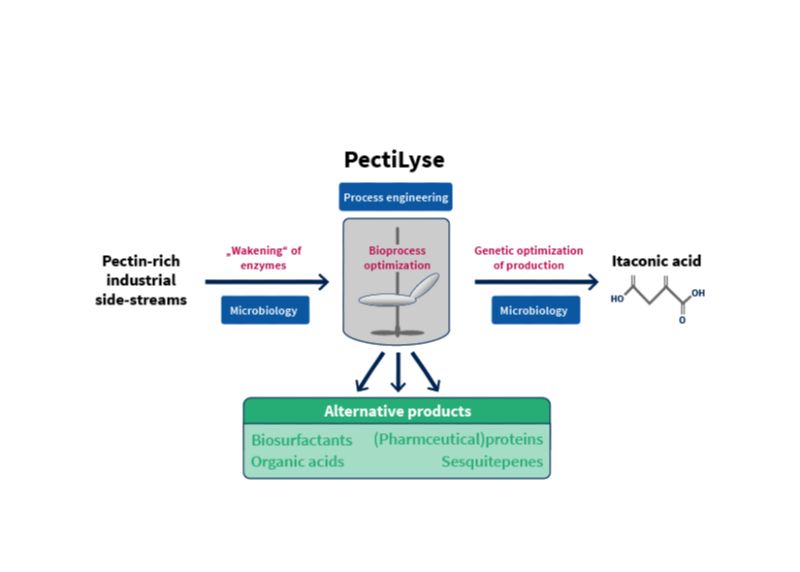
Enhancement of pectin-rich industrial side streams in a sustainable microbial bioprocess.
Copyright: Institute of Microbiology, HHU Düsseldorf
Its pathogenic lifestyle is reflected in the enzyme repertoire of Ustilago maydis. In contrast to other model fungi, such as baker´s yeast, Ustilago maydis already encodes enzymes to metabolize individual sugars contained in pectin. In addition, other enzymes useful for the degradation of pectin into these individual sugars are present. Unfortunately, these are synthesized only in the infectious phase during corn colonization. To solve this problem, the corresponding enzymes were kind of awakened and thus activated by molecular biological methods. Applying this “sleeping beauty” principle, the desired enzymes could successfully be produced in the yeast phase and secreted into the extracellular space. This allows an expansion of the substrate spectrum.
Break down a complex molecule
The main component of pectin is polygalacturonic acid, which forms the backbone of the complex molecule. The team thus first developed several Ustilago maydis strains that produce and secrete different enzymes for the degradation of polygalacturonic acid. For example, the researchers investigated the fungal enzyme endopolygalacturonase and assessed its effectivity. In addition, they transplanted already characterized, potent enzymes from other bacteria and fungi into their model organism Ustilago maydis by gene transfer. The result: the foreign enzymes were also produced successfully by this laboratory fungus and released into the culture medium without any problems. For production of the bacterial enzymes, the fact that Ustilago maydis has an unconventional secretion pathway that exports these enzymes without foreign modifications – with full activity potential – into the extracellular space was exploited.
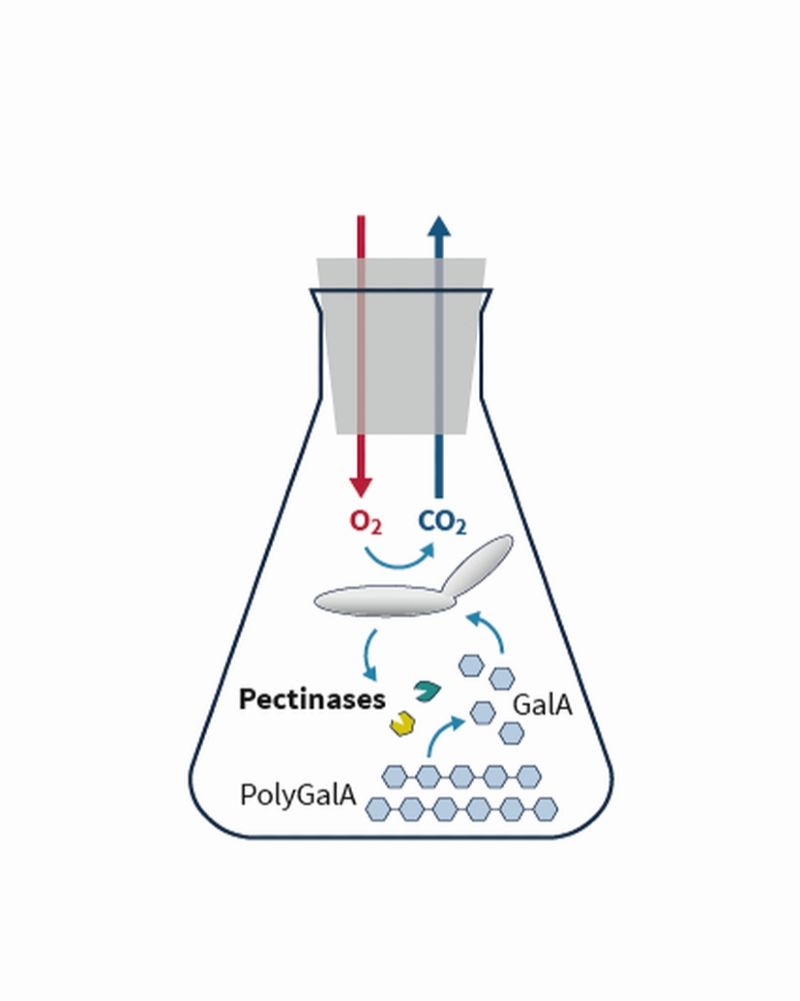
This broad approach allowed the identification of fungal strains that export different active pectinolytic enzymes. Using mixed cultures consisting of strains with complementary enzyme activities, efficient degradation of polygalacturonic acid (PolyGalA) to the component galacturonic acid (GalA), subsequently metabolized by the fungi, could finally be achieved. The project thus lays the foundation for further steps towards the degradation of pectin-containing biomass.
In order to successfully establish polygalacturonic acid degradation, modern online analytics from the biochemical engineers were key to evaluating the produced strains. The fungal strains were cultivated on polygalacturonic acid and, at the same time, their respiration rate was determined during the course of cultivation. Data analysis supported characterization of the individual strains with regard to the respective substrate turnover and enzyme activity. Indirect quantification of the polygalacturonic acid concentration could thus be achieved based on the respiration rate. Mixed cultures could also be analyzed efficiently in this way.
Further products in the focus
As an example of a valuable industrial product, the research team initially focused on the organic acid itaconate, which is naturally produced by Ustilago maydis under nitrogen limiting conditions. Itaconate is an important platform chemical in polymer chemistry. Its fields of application range from the production of paints and varnishes to thickeners for fats, pharmaceuticals or herbicides up to biologically degradable polymers. The biosynthesis of itaconate is regulated by a gene cluster and proved to be highly variable when compared in different Ustilago maydis strains. Very low itaconate production could be observed especially in the strain background used for secretion of pectin-degrading enzymes. New genetic approaches to remedy these variations could be developed by comparing biosynthetic enzymes of related smut fungi at the genome level. The team succeeded in permanently activating the central regulator of the gene cluster by a genetic trick. This turned out to be an excellent set screw for a high basic level of itaconate production. The method lays the foundation for the targeted bioprocess.
The PectiLyse project could thus establish the first basic steps on the way to an efficient bioprocess both at the level of substrate degradation as well as at the level of itaconate production. This was possible only by a team cooperating in the fields of microbiology, biotechnology and bioengineering. The researchers will further optimize the fungus for degradation of pectin in the future. At the same time, they are working on a consolidated bioprocess in which further valuable molecules are produced from the released fermentable sugars. In addition to itaconate, other products such as biosurfactants, organic acids, sesquiterpenes, which are used as flavorings or fragrances, or pharmaceutical proteins are conceivable. The production of biosurfactants was successfully established in the Bio2 BioSC
FocusLab. Based on the interdisciplinary work in the framework of the BioSC, Ustilago maydis could serve as a flexible platform organism for industrial biotechnology in the near future.
Project Coordinator
Dr. Kerstin Schipper
Institute of Microbiology, HHU Düsseldorf
Email
Participating Core Groups
Prof. Dr. Michael Feldbrügge, Dr. Kerstin Schipper, Dr. Benedikt Steuten, Dr. Peter Stoffels
Institute of Microbiology, HHU Düsseldorf
Prof. Dr. Lars M. Blank, Prof. Dr. Nick Wierckx1, Dr. Elena von Helden (née Geiser)
ABBt – Applied Microbiology, RWTH Aachen
Prof. Dr.-Ing. Jochen Büchs, Dr. Markus Müller, Dr. Tino Schlepütz
AVT – Bioprocess Engineering, RWTH Aachen
Funding Period
01.11.2015 – 31.10.2017
Funding
PectiLyse is part of the NRW-Strategieprojekt BioSC and thus funded by the Ministry of Culture and Science of the German State of North Rhine-Westphalia.
Publications
Geiser, E, Ludwig, F, Zambanini, T, Wierckx, N and Blank, LM (2016). Draft genome sequences of itaconate-producing ustilaginaceae. Genome Announc 4(6).
Geiser, E, Przybilla, SK, Engel, M, Kleineberg, W, Buttner, L, Sarikaya, E, Hartog, TD, Klankermayer, J, Leitner, W, Bolker, M, Blank, LM and Wierckx, N (2016). Genetic and biochemical insights into the itaconate pathway of ustilago maydis enable enhanced production. Metab Eng 38: 427-435.
Geiser, E, Reindl, M, Blank, LM, Feldbrugge, M, Wierckx, N and Schipper, K (2016). Activating intrinsic carbohydrate-active enzymes of the smut fungus ustilago maydis for the degradation of plant cell wall components. Appl Environ Microbiol 82(17): 5174-5185.
_______________________________________________
1 New address: IBG-1 Biotechnology, Forschungszentrum Jülich