Bioprocesses with microorganisms as biocatalysts play a decisive role in establishing a sustainable bioeconomy. In the framework of the BioSC project “Gluconobacter Factory”, an interdisciplinary research approach was used to explore the acetic acid bacterium Gluconobacter oxydans with regard to fundamental properties and application potentials. Among other findings, this resulted in a highly efficient production process for a potential new sweetener that is natural and low in calories.
Many biotechnological bioprocesses start with sugars such as glucose, fructose or sucrose. Microorganisms usually transport them initially into the interior of the cell, the cytoplasm, and then convert them via a variety of enzyme-catalyzed steps into the desired product, which then has to be exported from the cytoplasm. In Gluconobacter, however, numerous production processes take place outside the cytoplasm in the periplasm, which is the space between the cytoplasmic membrane and the outer membrane. Enzymes called dehydrogenases are located there, which oxidize sugars and other carbohydrates highly specifically at a specific carbon atom. The electrons released in the process are transferred via other enzymes (cytochrome oxidases) to oxygen, which is converted to water. Such specific oxidations are difficult or not at all possible to achieve purely chemically, so the superior capabilities of microorganisms are in demand. For these reasons, the bacterium Gluconobacter oxydans has already been used industrially for the production of vitamin C since the 1930s. In order to make even better and broader use of the biocatalytic potential of this bacterium, an interdisciplinary team worked on a detailed analysis of the genetic information and the development of new production strains and products.

Analysis of 2,664 genes
The genetic information of Gluconobacter oxydans in the form of deoxyribonucleic acid (DNA) is organized in a circular chromosome of 2.7 million base pairs with 2,432 genes and five plasmids with again a total of 232 genes. The question of how these more than 2,600 genes of Gluconobacter oxydans are organized into transcription units and which control sequences direct the transcription of DNA in ribonucleic acid (RNA) was answered by sequencing the RNA and subsequent bioinformatics analyses. This extensive information was made available to the scientific community on a freely accessible, web-based bioinformatics platform at www.gluconobacterfactory.de.
For the industrial use of Gluconobacter oxydans, large quantities of bacterial cells must first be cultivated. Here the special characteristics of Gluconobacter oxydans are disadvantageous: only about 100 g cells are formed from one kilogram of glucose, whereas for bacteria with a standard type of metabolism about 500 g cells are obtained. This means that the production of cells for biotransformations is considerably more expensive when Gluconobacter oxydans is used. By targeted modification of glucose metabolism, an approach called metabolic engineering, a strain gaining 160 g cells per kilogram of glucose could be constructed and thus allows reduction of the costs for production of cells.
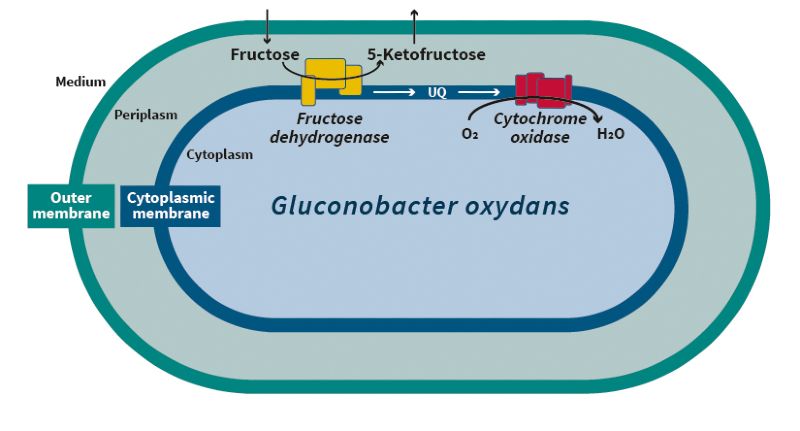
Production of a low-calorie sweetener
Due to its specific characteristics, Gluconobacter oxydans is highly suited for synthesizing membrane-bound, periplasmatic enzymes also from other bacteria in order to investigate their characteristics and application potentials. Fructose dehydrogenase from the related bacterium Gluconobacter japonicus proved to be especially promising. This enzyme consists of three different subunits and catalyzes the oxidation of fructose to 5-ketofructose. 5-ketofructose is a very interesting product since the compound has a comparable taste and a similar sweetening power as fructose itself, but unlike fructose, it seems to be not or only marginally metabolized by our intestinal flora. These are exactly the characteristics a low-calorie sweetener should have. The potential for the technical production of 5-ketofructose from fructose using Gluconobacter oxydans cells, which overproduce fructose dehydrogenase, was evaluated and optimized by the project team using methods of biochemical engineering. A process was developed that could produce 490 g of 5-ketofructose per liter with a product yield of 0.92 g per g fructose consumed and a space-time-yield of 7 g per liter and hour. This indicated that an economical production of the potential sweetener is possible. Further studies must now prove the tolerability of 5-ketofructose.
Patent application
The project shows that the concept of interdisciplinary cooperation in the framework of Boost Fund research projects is very successful and promising also in the future. It enables basic research as a prerequisite for innovations and the establishment of a circular bioeconomy based on renewable resources. The GLUFACT project resulted in a patent application and seven original publications in peer-reviewed journals. It was followed by two BMBF-funded projects (IMPRES und IMPRES-2), in which 5-ketofructose production was further developed and new products for human nutrition were established.
Prof. Dr. Michael Bott, Dr. Stephanie Bringer
IBG-1 Biotechnology, Forschungszentrum Jülich
E-Mail
Prof. Dr. Michael Bott, Dr. Stephanie Bringer, Dr. Ines Kiefler, Dr. Tino Polen, Dr. Angela Kranz
IBG-1 Biotechnology, Forschungszentrum Jülich
Prof. Dr. Björn Usadel, Dr. Alexander Vogel
IBG-4 Bioinformatics, Forschungszentrum Jülich
Prof. Dr. Uwe Deppenmeier, Dr. Konrad Kosciow, Dr. Anna Siemen
Institute for Microbiology and Biotechnology, Universität Bonn
Prof. Dr.-Ing. Jochen Büchs, Dr. Elena Herweg
AVT – Bioprocess Engineering, RWTH Aachen
1.1.2014 – 31.12.2016
GLUFACT is part of the NRW-Strategieprojekt BioSC and thus funded by the Ministry of Culture and Science of the German State of North Rhine-Westphalia.
Kiefler, I, Bringer, S and Bott, M (2015). Sdhe-dependent formation of a functional acetobacter pasteurianus succinate dehydrogenase in gluconobacter oxydans--a first step toward a complete tricarboxylic acid cycle. Appl Microbiol Biotechnol 99(21): 9147-9160.
Kosciow, K, Domin, C, Schweiger, P and Deppenmeier, U (2016). Extracellular targeting of an active endoxylanase by a tolb negative mutant of gluconobacter oxydans. J Ind Microbiol Biotechnol 43(7): 989-999.
Kosciow, K, Zahid, N, Schweiger, P and Deppenmeier, U (2014). Production of a periplasmic trehalase in gluconobacter oxydans and growth on trehalose. J Biotechnol 189: 27-35.
Loehrer, M, Vogel, A, Huettel, B, Reinhardt, R, Benes, V, Duplessis, S, Usadel, B and Schaffrath, U (2014). On the current status of phakopsora pachyrhizi genome sequencing. Front Plant Sci 5: 377.
Luchterhand, B, Fischoder, T, Grimm, AR, Wewetzer, S, Wunderlich, M, Schleputz, T and Buchs, J (2015). Quantifying the sensitivity of g. Oxydans atcc 621h and dsm 3504 to osmotic stress triggered by soluble buffers. J Ind Microbiol Biotechnol 42(4): 585-600.
Meyer, M, Schweiger, P and Deppenmeier, U (2015). Succinic semialdehyde reductase gox1801 from gluconobacter oxydans in comparison to other succinic semialdehyde-reducing enzymes. Appl Microbiol Biotechnol 99(9): 3929-3939.
Zahid, N and Deppenmeier, U (2016). Role of mannitol dehydrogenases in osmoprotection of gluconobacter oxydans. Appl Microbiol Biotechnol 100(23): 9967-9978.